
"This is a tsunami that’s moving right toward us.” That was the disquieting thought Stephen Streiffer, Argonne National Laboratory’s interim deputy lab director for science, had on January 10. That day, Chinese scientists first released the gene sequence of SARS-CoV-2, allowing researchers around the globe to begin study of the novel coronavirus. The very next day, China reported the first known death from COVID-19, the respiratory disease caused by the virus.
As the coronavirus began spreading throughout the world, infecting hundreds of thousands, Argonne’s top minds, along with associates from Northwestern University and the University of Chicago, joined forces on the front lines of the international scientific fight against the pandemic. At Argonne, a U.S. Department of Energy research center in Lemont, several teams of researchers — microbiologists, systems engineers, computer scientists, emergency management experts — have been feverishly working to provide answers to the most critical questions raised by the contagion, from how to create effective antiviral medications to which societal interventions will slow the spread of the disease and save the most lives. “The entirety of the infrastructure that we can bring to bear has got to be brought to bear to address this,” Streiffer says. “Otherwise people are going to keep dying.”
Here’s a breakdown of Argonne’s four main initiatives.

1. Know Your Enemy
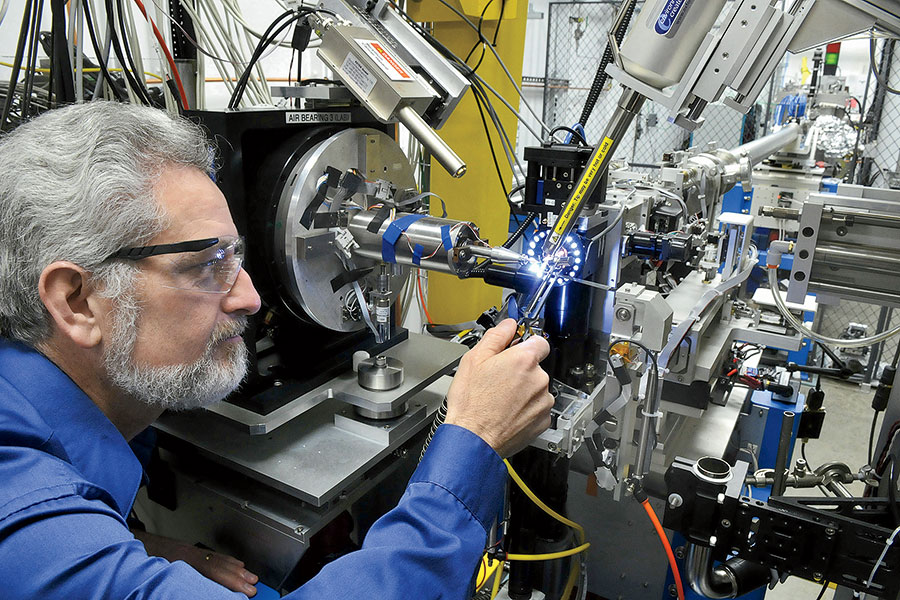
President Donald Trump has characterized the fight against the coronavirus as a war with an “invisible enemy.” But seeing the virus — and at Argonne, they can — is crucial to creating drugs to attack it. The lab’s Advanced Photon Source, an electron storage ring a kilometer in diameter, generates the brightest X-rays in the Western Hemisphere — a billion times more intense than those from the machine at your dentist’s office. Essentially functioning as a giant X-ray microscope, it allows researchers to study the virus at the atomic level.
Instead of risking infection by working with samples of the coronavirus, the researchers look at lab-grown crystals of its individual proteins. The crystals, each one about half the diameter of a human hair, are scooped onto a pin and blasted by an X-ray beam. “If you look at the accuracy with which we have to place a very, very tiny X-ray beam onto those crystals, it’s the equivalent of standing in Chicago and having a target sitting on the Space Needle in Seattle,” Streiffer says. “It’s so accurate that we have to account for the curvature of the earth.”
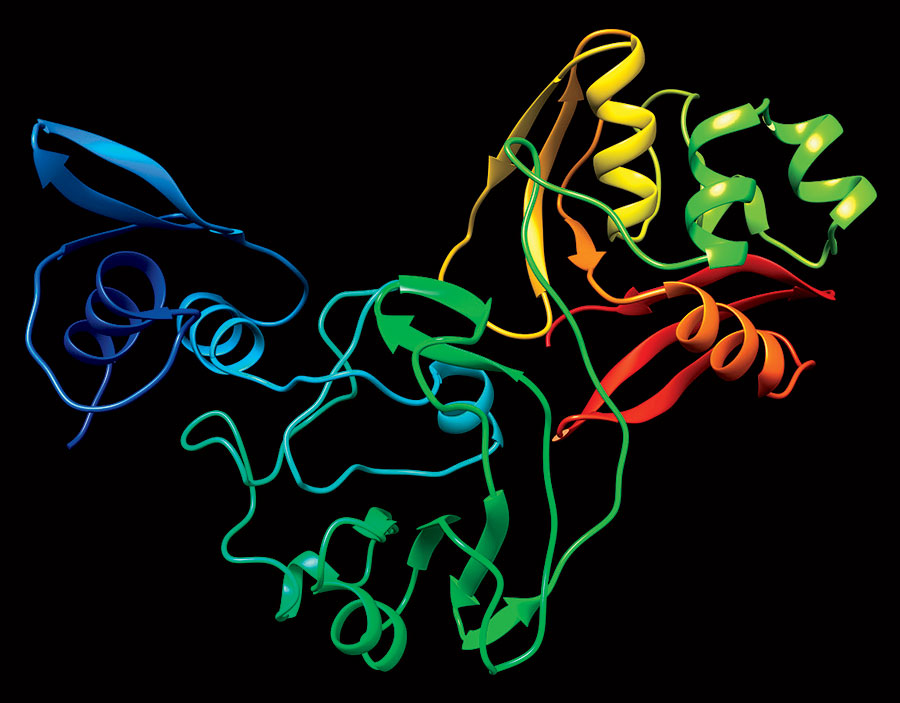
The APS yields 3D images of the proteins (they look like twisted ribbons), which are then cataloged in the Protein Data Bank, an online repository of biomolecular structures. While scientists intend on mapping all 28 coronavirus proteins, they pinpointed 15 as likely related to how the virus replicates in humans — an early step toward the creation of antiviral drugs. “Those are the proteins that have a greater probability that if you interrupt their function with some inhibitor, you may be able to stop the progression of the disease,” says Robert Fischetti, a leader of Argonne’s X-ray Science Division.
A team that includes Northwestern and U. of C. scientists made two key early discoveries. First they found a protein nearly identical to one in the virus that caused the 2002 SARS pandemic. Which means drugs that were in development before that disease was contained could now be seen to completion and used to treat COVID-19. They also located a protein complex within the virus that lets it hide from our immune system, allowing it time to replicate. The researchers believe a drug inhibiting that protein complex would slow down or stop the replication.
“I like to think of the protein as a machine,” says Karla Satchell, a professor of microbiology and immunology at Northwestern and a principal investigator for the Center for Structural Genomics of Infectious Diseases, whose team uses Argonne’s facilities. “You can throw a cog into the machine and keep it from working.”
2. Search and Destroy
Rick Stevens prefers a different analogy for an antiviral. “It’s like a lock and key: The lock is the virus protein, and the key is the drug,” says Stevens, Argonne’s associate laboratory director for computing, environment, and life sciences. “We’re trying to stick a key in and break it off in the lock.” The great unknown: In the vast universe of keys, which one fits this lock?
To find out, Stevens does the mother of all Google searches. Starting with a coronavirus protein structure derived from the APS, he and his colleagues comb through databases that hold information on some 10 billion molecules. The goal: identifying well-matched inhibitors that could lead to antiviral therapies. “The libraries include all known drugs,” Stevens says, “plus hundreds of millions of additional compounds that have been synthesized or could be synthesized.”
For these megasearches, several Argonne supercomputers work in concert with high-performance machines at other national laboratories. Each one, Stevens says, is “essentially a billion times faster than your cellphone.” With the aid of artificial intelligence, the system can find the needles in the haystack: substances that not only bond with a virus protein but are also safe for humans.
“We have hits of a few thousand molecules that we think are interesting,” Stevens says of a search related to an early target. His team narrows the results to a short list, which is passed to a structural biologist at Argonne, who then orders a lab to produce a crystal of the matching molecule bound to the virus protein. Next, APS researchers inspect the strength of the bonds.
Eventually, the information ends up in the hands of pharmaceutical companies and drug labs. But a lot of database sleuthing still remains to be done. “Drug development in the normal time frame takes years,” Stevens says. “We’re trying to do this in months.”
3. Theory of Evolution
Just as your family tree tells the story of your ancestry, the evolutionary tree of a virus tells the story of its origins, how it has changed over time, and how it spreads.
Using supercomputers, Argonne researchers are tracing the evolution of SARS-CoV-2 via its information-carrying unit: a single strand of RNA containing a gene sequence. “We have a few hundred samples from [infected people in] China, Europe, America, and the cruise ships,” Stevens explains. “All of those samples have been sequenced, and by comparing them, we can show how the virus has changed over the last few months.”
Like other single-strand RNA viruses, SARS-CoV-2 rapidly mutates. Researchers are closely monitoring those mutations to understand whether a virus that originated in an animal is continuing to adapt to humans, which proteins are changing in the process, and whether the virus has behaved differently over time, such as becoming deadlier or more contagious.
The good news on that grim front: “There is evidence in the past that when viruses become more host-adapted, they become less lethal,” Stevens explains. “From the virus’s standpoint, it might be advantageous not to kill the host so quickly or frequently to create more of a habitat. The only way we will understand that, as with everything in biology, is by putting it into comparative perspective through evolutionary analysis.”
4. The Mod Squad
The pandemic has had elected leaders and public health officials clamoring for solid information to guide policy decisions that have widespread life-and-death consequences. That’s where Argonne’s computer models step in to fill the information vacuum and help authorities devise a response that works — rather than, say, just hoping, as Trump once did, that “like a miracle, it will disappear.”
Using the laboratory’s supercomputers, researchers simulate the virus’s progression over time, dial up potential interventions, and predict the impact of those interventions on the infection rate. A single simulation can incorporate tens of thousands of parameters. Want to know how certain strategies — limiting gatherings, restricting travel, closing schools — will slow the rate of infection? Toggle them on and see. OK, now what if a third of the population doesn’t follow instructions? Click. In a worst-case scenario, what will the burden on the health care system be one month from now, or three? The simulation has the projection.
All of Argonne’s simulations — including one for a zombie apocalypse, which was released as something of a publicity stunt in 2016 — are modeled on Chicago. That’s because the research center’s “agent-based” framework, the Chicago Social Interaction Model, is built upon a mountain of local data. Researchers have now dubbed this model CityCOVID. Think of it as a scientifically true-to-life SimCity, populated by millions of individual “agents” with hour-by-hour activities — commuting, grocery shopping, going to school — that are statistically consistent with those of real Chicagoans. All the while, these avatars are potentially spreading disease. (Argonne adapted the model in 2014 for use during the spread of the Ebola virus in West Africa.)
“The key questions the model answers include: How many people will become infected? What is the peak and when will that occur? How do interventions affect these numbers, and over what time frame?” explains Charles Macal, who leads these simulations.
Macal’s team closely monitors the daily drip of scientific literature, finetuning the model with any new information. “How long does the virus incubate for? The CDC is continually working on it,” Macal says. “If they update their estimates, we’ll update the model.” Likewise, they’ve been keeping tabs on the shifting estimates of the COVID-19 mortality rate, from initial reports putting it around 2 percent, to the World Health Organization announcing in early March that it was 3.4 percent, to Anthony Fauci, director of the National Institute of Allergy and Infectious Diseases, more recently saying it’s “somewhere around 1 percent.”
City Hall and local hospitals are paying close attention to Argonne’s model, whose results Macal likens to a weather forecast. “If you know how many people are infected, you can apply a certain percentage of those that are probably serious or critical, and from that percentage you can predict how many health care workers you’ll need, how many hospital beds, how many ventilators, how much personal protective equipment.”
Macal ultimately hopes such information will save lives. And he looks forward to the day — many months from now, in all probability — that life begins to return to normal.


